- Home
- Member Resources
- Articles
- Tissue Provenance: Personalized Health Care Starts with Analyzing the Correct Person

Figure 1: Photomicrograph of unexpected tissue in a surgical pathology specimen.
Tissue provenance is the concept of assuring that tissue, whether in a container, on a slide or a swab, or in an instrument, is from the original source and authentic. Tissue "floaters" on slides (unexpected tissue types in a given case) or specimen mix-ups in pathology laboratories may confound anticipated histologic or other laboratory findings, complicating the already complex and difficult practice of anatomic pathology (Figure 1). Numerous reports over the past decades have examined this issue, finding unexpected tissues and/or genetic materials contaminating FFPE blocks, either at the time of embedding or in storage and transport, in up to 1 in 10 cases.1-5 While gross-dissection and histology laboratories strive to ensure high quality practices through cleanliness and regular equipment maintenance, encounters with floaters and specimen switches will arise at some point for most, if not all, pathologists. Access to identity testing may help to resolve these issues and can serve to reassure the pathologist dealing with a laboratory quality control issue or an unexpected finding, while ensuring that patients receive the proper diagnosis and appropriate follow up.
The University of Nebraska Medical Center (UNMC) Human DNA Identification Laboratory initially developed identity testing to support UNMC's bone marrow transplant program in the 1990s, shifting from CAP accreditation to ISO 17025:2017 accreditation to better align and support local law enforcement, providing testing for criminal justice purposes. ISO 17025:2017 accreditation is the standard for laboratories performing testing for forensic purposes. Over that time, Restriction Fragment Length Polymorphism (RFLP) analysis and subsequent Short Tandem Repeat (STR) analysis have resolved concerns surrounding specimen switches, suspected nonpatient tissue in anatomic pathology cases, and other specimen integrity concerns following environmental mishaps, for institutions around the country. In 2024, the UNMC laboratory performed testing for nearly 100 cases with a trend toward increasing volumes in 2025.
Many laboratories perform identity testing via STR profiling using commercially available reagents. STRs are repeating genomic elements distributed throughout the genome that can vary from person to person regarding repeat number. Interrogation of a selection of these elements are central to the determination of an individual’s "DNA-fingerprint." Most notably, these fingerprints are utilized for criminal justice purposes, as foundation to the Combined U.S. DNA Index System (CODIS) developed and maintained by the FBI.
The process of determining whether tissue from an individual other than the patient of record is tailored to the materials available. By convention in the UNMC laboratory, the suspect tissue is denoted as "unknown," and tissue expected to belong to the patient is denoted as "reference." This reference profile may originate from a separate, historical case known to belong to the patient, it may come from a buccal swab or blood draw, or it may come from the "uninvolved" tissue from the slide containing the tissue of concern.
For UNMC's laboratory testing, the initial approach to sampling depends heavily on the concern of the submitting client, the size of the tissue of concern, and the availability of materials to test. In cases where there is concern of pre-analytic error (eg, wrong tissue in wrong block), curls from FFPE blocks are taken from the block of concern and a separate block, known to be from the reference patient. For FFPE block-derived tissue floaters, submission of at least 5 unstained slides is ideal, in addition to the slide denoting the tissue of concern. For these unstained slides, H&E staining is performed, omitting coverslips, as this allows ready access to the tissue. The H&E staining helps to ensure that the cells of interest are definitively captured and gather a sense of how robust the isolation will be. For larger tissue fragments, sampling of the tissue can be done without magnification. For smaller cell clusters, cells are manually isolated with a scalpel and a dissection scope (Figure 2-C).
Certainly, this approach cannot always be used. At times, these tissues are fleeting, seen in deeper levels, only to be lost in subsequent specially stained slides (for immunohistochemistry, fungal stains, etc). In these instances, slides may be soaked in xylene, allowing for the removal of coverslips. In the UNMC experience, as little as 100 cells have provided usable STR locus information. Importantly, prior staining does not impair the ability to determine STR profiles. Regardless of whether the tissue sampled is stained or unstained, successful amplification of STR loci suitable for identification is obtainable. One important caveat for STR assays, and most any molecular technique, is decalcification of tissues using acidic solutions usually renders the specimen unusable. Acidic conditions lead to the depurination of DNA, thereby destroying templates needed for molecular analysis.
Following tissue sampling, the workflow and wet-bench work largely mirror standard forensics practices and procedures in the laboratory, with extracted DNA amplified by fluorescence multiplex PCR at STR loci and PCR products assessed using capillary electrophoresis. Upon availability of these STR profiles, peak positions may be compared between the unknown and reference sample, and conclusions may be drawn as to whether to tissue of concern originated from the same or different individuals.
Generally, this comparison is straight forward. The sampling techniques limit the co-mingling of tissues from reference and unknown, and if a two-person mixture is present, a definitive major contributor profile can be established. However, there have been identified instances where the unknown profile is close, but not an exact match when compared to the reference. In these instances, biology is important to remember. In most occasions, tissues of concern are often neoplastic cell clusters in an otherwise benign sample. Transformation of somatic cells is often driven by genetic mutation and other genomic instabilities. As STRs are genomic elements, they too are susceptible to the deficiencies present within tumor cells. As an example from the UNMC HLA Laboratory, a post-transplant chimerism study identified a mutated STR peak not seen previously in the pre-transplant samples following relapse of a patient's lymphoma (Figure 3).
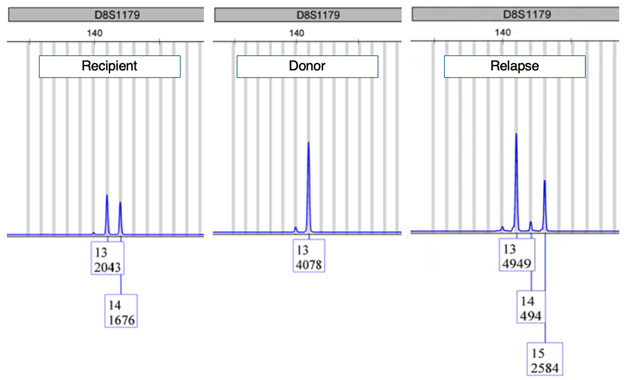
Figure 3: Electropherogram of the D8S1179 STR locus showing pretransplant profiles of recipient and donor prior to transplant and, following transplant with relapse of disease (B-cell lymphoma). The recipient locus with 14 repeats has expanded to 15 repeats in the tumor population.
Taken to a further extreme, the Human DNA Identification Laboratory recently tested a case where tumor was unexpectedly seen on the histomorphology slides following a seemingly normal endoscopy procedure. Identity testing was requested. The tissue cluster of concern was confirmed in deeper sections and carefully dissected and isolated from the surrounding uninvolved tissues. STR testing showed that the tallest STR peaks were shared with the patient’s reference sample; however, multiple STR loci showed a large number of additional peaks and a high degree of "stutter," an artifact commonly seen when amplifying repeating elements (Figure 4). Suspicious of microsatellite instability and/or mismatch repair (MMR) protein deficiency, MMR immunohistochemistry studies showed loss of MLH1 and PMS2 nuclear staining in the case, providing a biological explanation for the unanticipated STR peaks seen. In these circumstances, the concern for tissue floater was allayed and could be traced back to less-than-ideal documentation by the proceduralist.

Figure 4: Example of mismatch repair deficiency as seen on STR testing for identity confirmation. The "unknown" sample was isolated as a coherent cell population.
Identity testing can also be useful in sorting out potential confounding findings in molecular oncology assays. Contamination scores now accompany some NGS assays to assess for possible specimen contamination. As an example, the Nebraska Medicine clinical molecular oncology laboratory had received an outside FFPE block for large panel NGS somatic mutation testing. An initial H&E, several unstained slides (in case macrodissection was needed) and a final H&E were ordered. The unstained slides were scraped as macrodissection was not necessary for tumor enrichment. Upon NGS analysis, there was a high contamination score (with remaining specimens on the run having no issues and no variant patterns matching the contaminated specimen). Review of the initially cut H&E slide demonstrated no potentially contaminating tissue. After the high contamination score was identified, the final H&E was reviewed, which revealed a small piece of unexpected tissue that was confirmed by STR analysis to be foreign when compared to the reference. Thus, even in molecular testing, it is important to review all available H&E slides to verify no foreign tissue is present. Although unexpected and unrecognized foreign tissue could still exist in the block and affect molecular analysis, if the block is being extracted without cutting and macrodissection of unstained slides.
Tissue floaters and specimen mix-ups are a concern and a reality for the pathology community. Understanding methods for resolution, aside from assumption, can greatly reduce the stress and uncertainty for our anatomic pathologists, and ensure our patients receive the correct diagnosis and appropriate treatment and care.
References
- Platt E, Sommer P, McDonald L, Bennett A, Hunt J. Tissue floaters and contaminants in the histology laboratory. Arch Pathol Lab Med. 2009;133(6):973-978.
- Lamothe S, Peric M, Glickman JN, Heher YK. Erroneous patient tissue contaminants in 1574 surgical pathology slides: impact on diagnostic error and a novel framework for floater management. Arch Pathol Lab Med. 2023;147(12):1413-1421.
- Carll T, Fuja C, Antic T, Lastra R, Pytel P. Tissue contamination during transportation of formalin-fixed, paraffin-embedded blocks. Am J Clin Pathol. 2022;158(1):96-104.
- Sapp AR, Wander P, Ali U, Gong Y, Milano RV, Bartel MJ. Discordant pathological and endoscopic diagnosis: consider floaters. ACG Case Rep J. 2021;8(5):e00583.
- Hodgson AJ, Shang YM, Boulianne P, Downes MR, Hwang D, Slodkowska E. A practical approach to investigating cross-contaminants in the anatomical pathology laboratory. Int J Surg Pathol. 2020;28(7):700-710.

Jesse Cox, MD, PhD, FCAP, is an Associate Professor and Molecular Pathologist in the Department of Pathology Microbiology and Immunology at the University of Nebraska Medical Center in Omaha, Nebraska. Dr. Cox serves as medical director of the Human DNA Identification Laboratory, as well as director of the UNMC HLA Laboratory, Nebraska Public Service Laboratory and Pathology Residency Training Program. He also serves significant roles in Molecular Diagnostics and other Clinical Pathology areas.

Allison Cushman-Vokoun, MD, PhD, FCAP, is the medical director of both the Molecular Diagnostics Laboratory and the Warren G. Sanger Human Genetics Laboratory at the University of Nebraska Medical Center/Nebraska Medicine in Omaha, Nebraska. She is one of the leaders for the Precision Medicine Initiative at Nebraska Medicine, and she started the Molecular Genetic Pathology Fellowship Program at UNMC. She is a full-time molecular pathologist with research interests in the translational evaluation of molecular biomarkers in various malignancies. She has happily served on two terms of the CAP Personalized Health Care Committee.


